CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Glycosyltransferases"
| Line 38: | Line 38: | ||
|- | |- | ||
| − | !style="width: | + | !style="width:33%"|A representative GT-A fold: SpsA from ''Bacillus subtilus'', PDB code [{{PDBlink}}1h7l 1h7l] <cite>Charnock1999</cite>. The complex also contains two magnesium ions and a molecule of thymidine-5'-diphosphate. |
| − | !style="width: | + | !style="width:33%"|A representative GT-B fold: beta-glucosyltransferase from bacteriophage T4, PDB code [{{PDBlink}}1bgu 1bgu] <cite>Vrielink1994</cite>. The complex also contains a molecule of uridine-5'-diphosphate. |
|- | |- | ||
| | | | ||
Revision as of 15:41, 15 August 2010
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: Spencer Williams
- Responsible Curator: Spencer Williams
Overview
Glycosyltransferases are enzymes that catalyze the formation of the glycosidic linkage to form a glycoside. These enzymes utilize 'activated' sugar phosphates as glycosyl donors, and catalyze glycosyl group transfer to a nucleophilic group, usually an alcohol. The product of glycosyl transfer may be an O-, N-, S-, or C-glycoside; the glycoside may be part of a monosaccharide glycoside, oligosaccharide, or polysaccharide ([1, 2, 3, 4, 5]).
Donors
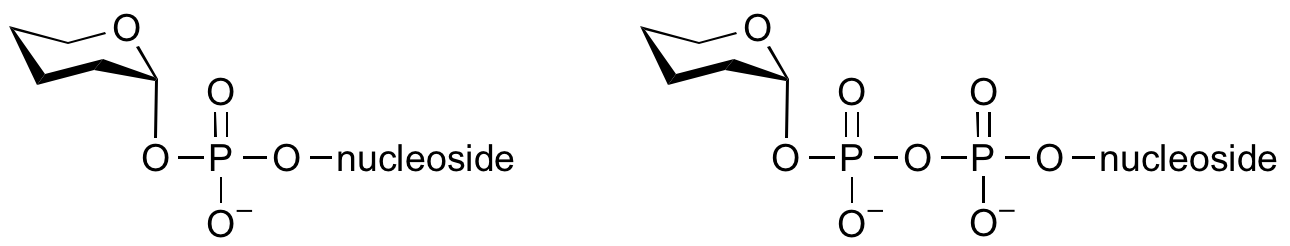
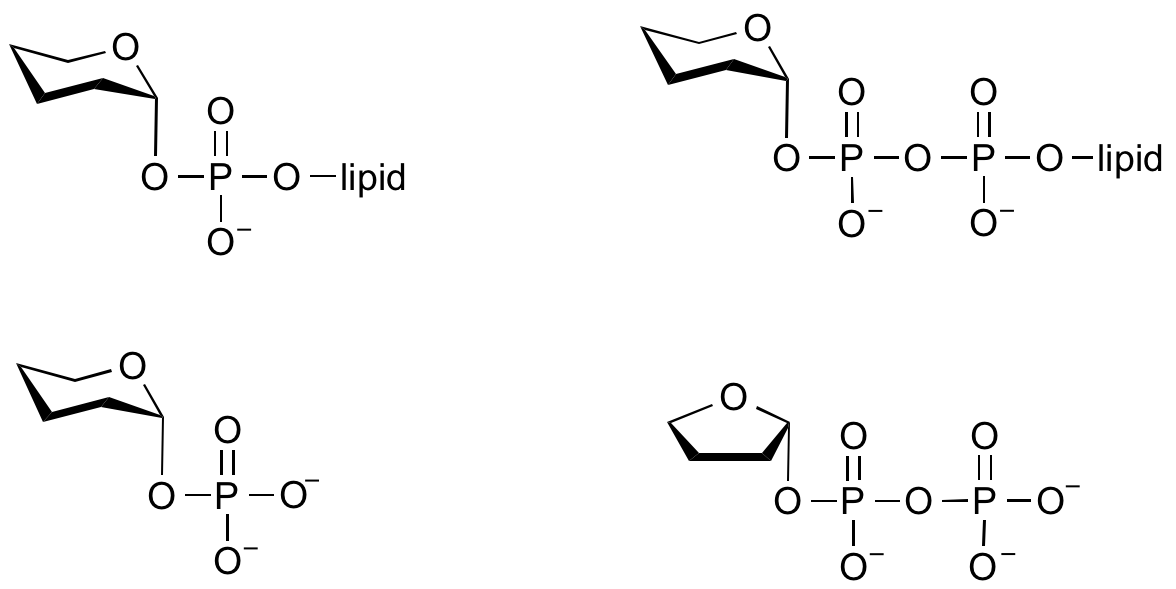
Glycosyltransferases can utilize a range of donor species. Sugar mono- or diphosphonucleotides are sometimes termed Leloir donors (after Nobel prize winner, Luis Leloir); the corresponding enzymes are termed Leloir donors.
Glycosyltransferases that utilize non-nucleotide donors, which may be polyprenol pyrophosphates, polyprenol phosphates, sugar-1-phosphates, or sugar-1-pyrophosphates, are termed non-Leloir glycosyltransferases.
Mechanism
Glycosyltransferases catalyze the transfer of glycosyl groups to a nucleophilic acceptor with either retention or inversion of configuration at the anomeric centre. This allows the classification of glycosyltransferases as either retaining or inverting enzymes.
Inverting glycosyltransferases
Structural and kinetic data for inverting glycosyltransferases support a mechanism that proceeds through a single nucleophilic substitution step, facilitated by an enzymic general base catalyst. The transition state is believed to possess substantial oxocarbenium ion character.
Retaining glycosyltransferases
Classification
Sequence based classification
Sequence-based classification uses algorithmic methods to assign sequences to various families. The glycosyltransferases have been classified into more than 90 families [6, 7]; this is permanently available through the Carbohydrate Active enZyme database [8]. Each family (GT family) contains proteins that are related by sequence, and by corollary, fold. This allows several useful predictions to be made since the catalytic machinery is conserved within each family. Usually, the mechanism used (ie retaining or inverting) is conserved within a GT family.
3-D folds
Leloir GTs
Sugar nucleotide-dependent (Leloir) glycosyltransferases have been found to possess only two different folds, termed the GT-A and GT-B folds. The GT-A fold is typified by the first member to have its X-ray structure determined, SpsA fromBacillus subtilus [9]. The GT-B fold was exemplified by its first member, the beta-glucosyltransferase from bacteriophage T4 [10].
Example structures
| A representative GT-A fold: SpsA from Bacillus subtilus, PDB code 1h7l [9]. The complex also contains two magnesium ions and a molecule of thymidine-5'-diphosphate. | A representative GT-B fold: beta-glucosyltransferase from bacteriophage T4, PDB code 1bgu [10]. The complex also contains a molecule of uridine-5'-diphosphate. |
|---|---|
|
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1h7l.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill;set spin Y 10; spin on; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1BGU.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill;set spin Y 10; spin on; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
Non-Leloir GTs
Non-Leloir glycosyltransferases possess other folds. For example the polymerizing glycosyltransferase transglycosylase, which catalyzes the condensation of oligosaccharyl polyprenolphosphate to generate the carbohydrate backbone of peptidoglycan has a 'lysozyme'-like fold....
Example structures
| A representative GT-A fold: SpsA from Bacillus subtilus, PDB code 1h7l [9]. The complex also contains two magnesium ions and a molecule of thymidine-5'-diphosphate. | A representative GT-B fold: beta-glucosyltransferase from bacteriophage T4, PDB code 1bgu [10]. |
|---|---|
|
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1h7l.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill;set spin Y 10; spin on; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>1BGU.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill;set spin Y 10; spin on; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
Role of metals
Common sugar nucleotide donors
References
- Lairson LL, Henrissat B, Davies GJ, and Withers SG. (2008). Glycosyltransferases: structures, functions, and mechanisms. Annu Rev Biochem. 2008;77:521-55. DOI:10.1146/annurev.biochem.76.061005.092322 |
- Coutinho PM, Deleury E, Davies GJ, and Henrissat B. (2003). An evolving hierarchical family classification for glycosyltransferases. J Mol Biol. 2003;328(2):307-17. DOI:10.1016/s0022-2836(03)00307-3 |
Chapter 5: Coutinho PM, Rancurel C, Stam M, Bernard T, Couto FM, Danchin EGJ, Henrissat B. "Carbohydrate-active Enzymes Database: Principles and Classification of Glycosyltransferases."
- Campbell JA, Davies GJ, Bulone V, and Henrissat B. (1997). A classification of nucleotide-diphospho-sugar glycosyltransferases based on amino acid sequence similarities. Biochem J. 1997;326 ( Pt 3)(Pt 3):929-39. DOI:10.1042/bj3260929u |
-
Carbohydrate Active Enzymes database; URL http://www.cazy.org/
- Charnock SJ and Davies GJ. (1999). Structure of the nucleotide-diphospho-sugar transferase, SpsA from Bacillus subtilis, in native and nucleotide-complexed forms. Biochemistry. 1999;38(20):6380-5. DOI:10.1021/bi990270y |
- Vrielink A, Rüger W, Driessen HP, and Freemont PS. (1994). Crystal structure of the DNA modifying enzyme beta-glucosyltransferase in the presence and absence of the substrate uridine diphosphoglucose. EMBO J. 1994;13(15):3413-22. DOI:10.1002/j.1460-2075.1994.tb06646.x |