CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Glycoside Hydrolase Family 99"
| Line 29: | Line 29: | ||
== Substrate specificities == | == Substrate specificities == | ||
| + | [[Image:GH99_substrates.png|thumb|right|300px|'''Figure 1. Endo-α-mannosidase is located in the Golgi apparatus and pre-Golgi intermediates and acts on folded and unfolded ER-escaped glucosylated N-linked glycoproteins (X = protein); glucosylated free oligosaccharides (X = OH); and dolichol-bound glucosylated glycans (X = diphosphodolichol). Endo-α-mannosidase is an endo-acting glycoside hydrolase that cleaves the glycosidic bond between the glucose-substituted mannose and the rest of the glycan (see arrow). Endo-α-mannosidase substrates may arise from: (a) GI/GII inhibitor blockade or genetic knockout. (b) Glycoproteins that escape the CNX/CRT cycle without GII cleavage that are trimmed by ER and/or Golgi α-mannosidases. Endo-α-mannosidase also appears to act on glucosylated Man8GlcNAc and Man7GlcNAc, glycan structures that are poor substrates for GII; and (c) free oligosaccharides.''']] | ||
| + | |||
| + | |||
[[Glycoside hydrolases]] of family GH99 are [[endo]]-acting α-mannosidases that cleave glucose-substituted mannose within immature N-linked glycans of the general formula Glc<sub>1-3</sub>Man<sub>9</sub>GlcNAc<sub>2</sub> (or structures trimmed in the B and C branches), and possess maximal activity on the monoglucosylated forms <cite>Roth2003</cite>. This family was originally created from the mammalian enzyme, cloned by Spiro and co-workers <cite>Spiro1997</cite>. Mammalian GH99 enzymes are localized to the Golgi apparatus <cite>Zuber2000</cite> and appear to play a role in the rescue of glucosylated N-linked glycans that have evaded the action of the endoplasmic reticulum ''exo''-glucosidases I and II <cite>Dale1986</cite>. Mammalian [[endo]]-α-mannosidase has greater activity on glucosylated N-linked glycans that have been trimmed in the non-glucose-substituted branches <cite>Spiro1997</cite>. There is evidence that mammalian [[endo]]-α-mannosidases act on dolichol-bound N-glycan precursors <cite>Spiro2000</cite>, as well as free oligosaccharides released from N-glycoproteins and which undergo retrograde transport through the secretory pathway <cite>Kukushkin2011</cite>. Substrate studies on a bacterial GH99 enzyme from ''Shewanella amazonensis'' <cite>Matsuda2011</cite>, and the ''Bacteroides thetaiotaomicron'' and ''Bacteroides xylanisolvens'' enzymes <cite>Thompson2012</cite> have shown that these enzymes can also process Glc<sub>1/3</sub>Man<sub>9/7</sub>GlcNAc<sub>2</sub> structures, matching the substrate specificity of the mammalian enzymes. Kinetic analysis of ''Bacteroides thetaiotaomicron'' GH99 on a fluorescently-labelled Glc<sub>3</sub>Man<sub>9</sub>GlcNAc<sub>2</sub> structure yielded kinetic parameters that were similar to that found for the mammalian enzyme <cite>Thompson2012</cite>. Both mammalian and bacterial enzymes can utilize simple 'reducing end' substrate mimics such as methylumbelliferyl α-glucosyl-1,3-α-mannopyranoside <cite>Vogel2000</cite> or α-glucosyl-1,3-α-mannopyranosyl fluoride <cite>Thompson2012</cite>, but are inactive on aryl mannosides. | [[Glycoside hydrolases]] of family GH99 are [[endo]]-acting α-mannosidases that cleave glucose-substituted mannose within immature N-linked glycans of the general formula Glc<sub>1-3</sub>Man<sub>9</sub>GlcNAc<sub>2</sub> (or structures trimmed in the B and C branches), and possess maximal activity on the monoglucosylated forms <cite>Roth2003</cite>. This family was originally created from the mammalian enzyme, cloned by Spiro and co-workers <cite>Spiro1997</cite>. Mammalian GH99 enzymes are localized to the Golgi apparatus <cite>Zuber2000</cite> and appear to play a role in the rescue of glucosylated N-linked glycans that have evaded the action of the endoplasmic reticulum ''exo''-glucosidases I and II <cite>Dale1986</cite>. Mammalian [[endo]]-α-mannosidase has greater activity on glucosylated N-linked glycans that have been trimmed in the non-glucose-substituted branches <cite>Spiro1997</cite>. There is evidence that mammalian [[endo]]-α-mannosidases act on dolichol-bound N-glycan precursors <cite>Spiro2000</cite>, as well as free oligosaccharides released from N-glycoproteins and which undergo retrograde transport through the secretory pathway <cite>Kukushkin2011</cite>. Substrate studies on a bacterial GH99 enzyme from ''Shewanella amazonensis'' <cite>Matsuda2011</cite>, and the ''Bacteroides thetaiotaomicron'' and ''Bacteroides xylanisolvens'' enzymes <cite>Thompson2012</cite> have shown that these enzymes can also process Glc<sub>1/3</sub>Man<sub>9/7</sub>GlcNAc<sub>2</sub> structures, matching the substrate specificity of the mammalian enzymes. Kinetic analysis of ''Bacteroides thetaiotaomicron'' GH99 on a fluorescently-labelled Glc<sub>3</sub>Man<sub>9</sub>GlcNAc<sub>2</sub> structure yielded kinetic parameters that were similar to that found for the mammalian enzyme <cite>Thompson2012</cite>. Both mammalian and bacterial enzymes can utilize simple 'reducing end' substrate mimics such as methylumbelliferyl α-glucosyl-1,3-α-mannopyranoside <cite>Vogel2000</cite> or α-glucosyl-1,3-α-mannopyranosyl fluoride <cite>Thompson2012</cite>, but are inactive on aryl mannosides. | ||
| − | + | ||
== Kinetics and Mechanism == | == Kinetics and Mechanism == | ||
Family GH99 [[endo]]-α-mannosidases are [[retaining]] enzymes, as first shown by <sup>1</sup>H NMR analysis of the hydrolysis of α-glucosyl-1,3-α-mannopyranosyl fluoride by ''Bacteroides thetaiotaomicron'' <cite>Thompson2012</cite>. [[Retaining]] mannoside hydrolases (eg [[GH38]]) typically follow a [[classical Koshland double-displacement mechanism]]. However, X-ray crystallographic analysis of ''Bx''GH99 and ''Bt''GH99 failed to reveal a candidate nucleophilic residue; instead an alternative mechanism involving substrate-assisted catalysis by the 2OH residue and proceeding through a 1,2-anhydro sugar was proposed <cite>Thompson2012</cite>. In this proposal, Glu333 in ''BxGH99'' (Glu329 in ''Bt''GH99) acts as a [[general acid/base]] to deprotonate the 2OH and protonate the 1,2-anhydrosugar. | Family GH99 [[endo]]-α-mannosidases are [[retaining]] enzymes, as first shown by <sup>1</sup>H NMR analysis of the hydrolysis of α-glucosyl-1,3-α-mannopyranosyl fluoride by ''Bacteroides thetaiotaomicron'' <cite>Thompson2012</cite>. [[Retaining]] mannoside hydrolases (eg [[GH38]]) typically follow a [[classical Koshland double-displacement mechanism]]. However, X-ray crystallographic analysis of ''Bx''GH99 and ''Bt''GH99 failed to reveal a candidate nucleophilic residue; instead an alternative mechanism involving substrate-assisted catalysis by the 2OH residue and proceeding through a 1,2-anhydro sugar was proposed <cite>Thompson2012</cite>. In this proposal, Glu333 in ''BxGH99'' (Glu329 in ''Bt''GH99) acts as a [[general acid/base]] to deprotonate the 2OH and protonate the 1,2-anhydrosugar. | ||
| − | [[Image: | + | [[Image:GH99_mechanisms.png|thumb|center|900px|'''Figure 2. Proposed catalytic mechanisms for GH99 endo-α-mannosidase. (A) [[Classical Koshland double-displacement mechanism]]. (B) Neighboring group participation via a 1,2-anhydrosugar intermediate.''']] |
== Catalytic Residues == | == Catalytic Residues == | ||
Revision as of 18:21, 9 January 2012
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Spencer Williams^^^
- Responsible Curator: ^^^Gideon Davies^^^
| Glycoside Hydrolase Family GH99 | |
| Clan | not assigned |
| Mechanism | retaining |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/GH99.html | |
Substrate specificities
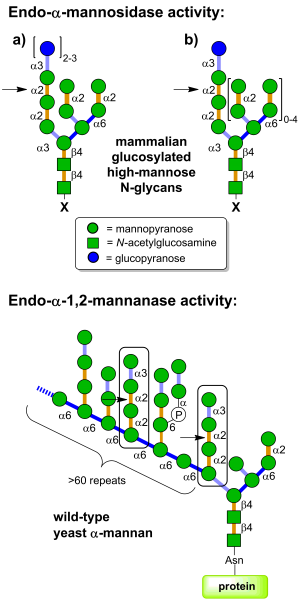
Glycoside hydrolases of family GH99 are endo-acting α-mannosidases that cleave glucose-substituted mannose within immature N-linked glycans of the general formula Glc1-3Man9GlcNAc2 (or structures trimmed in the B and C branches), and possess maximal activity on the monoglucosylated forms [1]. This family was originally created from the mammalian enzyme, cloned by Spiro and co-workers [2]. Mammalian GH99 enzymes are localized to the Golgi apparatus [3] and appear to play a role in the rescue of glucosylated N-linked glycans that have evaded the action of the endoplasmic reticulum exo-glucosidases I and II [4]. Mammalian endo-α-mannosidase has greater activity on glucosylated N-linked glycans that have been trimmed in the non-glucose-substituted branches [2]. There is evidence that mammalian endo-α-mannosidases act on dolichol-bound N-glycan precursors [5], as well as free oligosaccharides released from N-glycoproteins and which undergo retrograde transport through the secretory pathway [6]. Substrate studies on a bacterial GH99 enzyme from Shewanella amazonensis [7], and the Bacteroides thetaiotaomicron and Bacteroides xylanisolvens enzymes [8] have shown that these enzymes can also process Glc1/3Man9/7GlcNAc2 structures, matching the substrate specificity of the mammalian enzymes. Kinetic analysis of Bacteroides thetaiotaomicron GH99 on a fluorescently-labelled Glc3Man9GlcNAc2 structure yielded kinetic parameters that were similar to that found for the mammalian enzyme [8]. Both mammalian and bacterial enzymes can utilize simple 'reducing end' substrate mimics such as methylumbelliferyl α-glucosyl-1,3-α-mannopyranoside [9] or α-glucosyl-1,3-α-mannopyranosyl fluoride [8], but are inactive on aryl mannosides.
Kinetics and Mechanism
Family GH99 endo-α-mannosidases are retaining enzymes, as first shown by 1H NMR analysis of the hydrolysis of α-glucosyl-1,3-α-mannopyranosyl fluoride by Bacteroides thetaiotaomicron [8]. Retaining mannoside hydrolases (eg GH38) typically follow a classical Koshland double-displacement mechanism. However, X-ray crystallographic analysis of BxGH99 and BtGH99 failed to reveal a candidate nucleophilic residue; instead an alternative mechanism involving substrate-assisted catalysis by the 2OH residue and proceeding through a 1,2-anhydro sugar was proposed [8]. In this proposal, Glu333 in BxGH99 (Glu329 in BtGH99) acts as a general acid/base to deprotonate the 2OH and protonate the 1,2-anhydrosugar.

Catalytic Residues
The general acid/base was highlighted by X-ray crystallographic analysis as Glu336 in BxGH99 and Glu332 in BtGH99 [8]. The Glu332Ala mutant of BtGH99 shows a 50-fold decrease in catalytic activity relative to the wild-type enzyme using the activated substrate α-glucosyl-1,3-α-mannopyranosyl fluoride, and undetectable activity against the natural substrate, Glc3Man7GlcNAc2, supporting the role of this residue as general acid/base. As described in "Kinetics and Mechanism" the identity of the nucleophilic residue used for catalysis is more obscure and the 2OH of the substrate has been proposed to act as a nucleophile in a mechanism involving substrate assisted catalysis [8]. According to this proposal, Glu333 in BxGH99 (Glu329 in BtGH99) acts as a general acid/base to deprotonate the 2OH and protonate the 1,2-anhydrosugar.
Three-dimensional structures
Three-dimensional structures are available for bacterial members of GH99, including Bacteroides thetaiotaomicron and Bacteroides xylanisolvens [8]. They have a classical (β/α)8 TIM barrel fold, which is distinguished by the presence of extended loop motifs that form the active site. In different structures of the bacterial enzymes, these loops adopt different conformations, and appear to play a role in recognizing the extended structure of the N-glycan substrate. Binary complexes with two inhibitors, α-glucosyl-1,3-deoxymannonojirimycin and α-glucosyl-1,3-isofagomine, and 'active-site-spanning' ternary complexes with the same two inhibitors and the reducing end product fragment 1,2-α-mannobiose, provided insight into active site residues, especially the acid/base (Glu336 in BxGH99; Glu332 in BtGH99) and another key residue that interacts with both the 2OH of the -1 mannose residue and the 6OH of the -2 glucose residue, which provides a rationale for the requirement of a glucosylated-mannoside as the minimal substrate for GH99 enzymes [8]. As discussed in more detail in the "Kinetics and Mechanism" section, the precise identity of the nucleophilic residue is unclear, as in all GH99-inhibitor complexes with an occupied -1 subsite there is no candidate nucleophile within a reasonable distance to the "anomeric" carbon: in BxGH99 Glu333 is approximately 3.5 Å distant and the OH of Tyr46 and Tyr252 are 4.0 Å distant.
Sample structures
| Three-dimensional structure of GH99 endo-α-mannosidase from Bacteroides xylanisolvens, PDB code [1] [8]. | Three-dimensional structure of GH99 endo-α-mannosidase from Bacteroides xylanisolvens bound to glucose-1,3-isofagomine and α-1,2- mannobiose, PDB code [2] [8]. |
|---|---|
|
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>4ad1.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select ligand; wireframe 0.3; select MG; spacefill; set spin Y 10; spin off; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
<jmol> <jmolApplet> <color>white</color> <frame>true</frame> <uploadedFileContents>4ad4.pdb</uploadedFileContents> <script>cpk off; wireframe off; cartoon; color cartoon powderblue; select GLC or IFG; wireframe 0.3; select MAN; wireframe 0.3; set spin Y 10; spin off; set antialiasDisplay OFF</script> </jmolApplet> </jmol> |
Family Firsts
- First sterochemistry determination
- Bacteroides thetaiotaomicron endo-α-mannosidase by 1H NMR [8]
- First catalytic nucleophile identification
- It has been proposed that GH99 enzymes operate through a mechanism involving substrate assisted catalysis by the 2OH group of the -1 mannose residue [8]
- First general acid/base residue identification
- Bacteroides thetaiotaomicron endo-α-mannosidase by X-ray crystallography and analysis of enzyme mutant activities [8]
- First 3-D structure of a GH99 enzyme
- Bacteroides thetaiotaomicron and Bacteroides xylanisolvens endo-α-mannosidases [8]
References
- Roth J, Ziak M, and Zuber C. (2003). The role of glucosidase II and endomannosidase in glucose trimming of asparagine-linked oligosaccharides. Biochimie. 2003;85(3-4):287-94. DOI:10.1016/s0300-9084(03)00049-x |
- Spiro MJ, Bhoyroo VD, and Spiro RG. (1997). Molecular cloning and expression of rat liver endo-alpha-mannosidase, an N-linked oligosaccharide processing enzyme. J Biol Chem. 1997;272(46):29356-63. DOI:10.1074/jbc.272.46.29356 |
- Zuber C, Spiro MJ, Guhl B, Spiro RG, and Roth J. (2000). Golgi apparatus immunolocalization of endomannosidase suggests post-endoplasmic reticulum glucose trimming: implications for quality control. Mol Biol Cell. 2000;11(12):4227-40. DOI:10.1091/mbc.11.12.4227 |
- Dale MP, Kopfler WP, Chait I, and Byers LD. (1986). Beta-glucosidase: substrate, solvent, and viscosity variation as probes of the rate-limiting steps. Biochemistry. 1986;25(9):2522-9. DOI:10.1021/bi00357a036 |
- Spiro MJ and Spiro RG. (2000). Use of recombinant endomannosidase for evaluation of the processing of N-linked oligosaccharides of glycoproteins and their oligosaccharide-lipid precursors. Glycobiology. 2000;10(5):521-9. DOI:10.1093/glycob/10.5.521 |
- Kukushkin NV, Alonzi DS, Dwek RA, and Butters TD. (2011). Demonstration that endoplasmic reticulum-associated degradation of glycoproteins can occur downstream of processing by endomannosidase. Biochem J. 2011;438(1):133-42. DOI:10.1042/BJ20110186 |
-
Matsuda K, Kurakata Y, Miyazaki T, Matsuo I, Ito Y, Nishikawa A, Tonozuka T. Heterologous expression, purification, and characterization of an α-mannosidase belonging to glycoside hydrolase family 99 of Shewanella amazonensis. Biosci Biotechnol Biochem. 2011;75(4):797-9.
Note: Due to a problem with PubMed data, this reference is not automatically formatted. Please see these links out: DOI: 10.1271/bbb.100874 PMID:21512220
-
Thompson AJ, Williams RJ, Hakki Z, Alonzi DS, Wennekes T, Gloster TM, Songsrirote K, Thomas-Oates JE, Stütz AE, Butters TD, Williams SJ, Davies GJ. Structural and mechanistic insight into N-glycan processing by endo-α-mannosidase, Proc. Natl. Acad. Sci. USA., in press.
Note: Due to a problem with PubMed data, this reference is not automatically formatted. Please see these links out: DOI: 10.1073/pnas.1111482109 PMID:22219371
-
Vogel C, Pohlentz G. Synthesis of α-D-glucopyranosyl-(1,3)-α-D-mannopyranosyl-(1,7)-4-methylumbelliferone, a fluorogenic substrate for endo-α-1,2-mannosidase. J. Carbohydr. Chem. 2000, 19: 1247-1258.