CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Glycoside Hydrolase Family 162"
| Line 56: | Line 56: | ||
== Catalytic Residues == | == Catalytic Residues == | ||
The general acid and base of ''Tf''SGL are E262 and D446, respectively <cite>Tanaka2019</cite>. Both residues are highly conserved in [[GH162]] enzymes. The general acid of ''Tf''SGL is well superimposed with an acidic residue in a [[GH144]] bacterial β-1,2-glucanase from ''Chitinophaga pinensis'' (''Cp''SGL), whereas the general base is not superimposed <cite>Tanaka2019, Abe2017</cite>. Although the reaction mechanisms of [[GH144]] are unclear as of June 4th, 2019, ''Tf''SGL is clearly different from [[GH144]] in the reaction mechanism based on structural comparison <cite>Tanaka2019</cite>. | The general acid and base of ''Tf''SGL are E262 and D446, respectively <cite>Tanaka2019</cite>. Both residues are highly conserved in [[GH162]] enzymes. The general acid of ''Tf''SGL is well superimposed with an acidic residue in a [[GH144]] bacterial β-1,2-glucanase from ''Chitinophaga pinensis'' (''Cp''SGL), whereas the general base is not superimposed <cite>Tanaka2019, Abe2017</cite>. Although the reaction mechanisms of [[GH144]] are unclear as of June 4th, 2019, ''Tf''SGL is clearly different from [[GH144]] in the reaction mechanism based on structural comparison <cite>Tanaka2019</cite>. | ||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
| − | |||
Revision as of 00:53, 5 June 2019
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Nobukiyo Tanaka^^^
- Responsible Curator: ^^^Masahiro Nakajima^^^
| Glycoside Hydrolase Family GH162 | |
| Clan | GH-x |
| Mechanism | Inverting |
| Active site residues | Known |
| CAZy DB link | |
| https://www.cazy.org/GH162.html | |
Substrate specificities
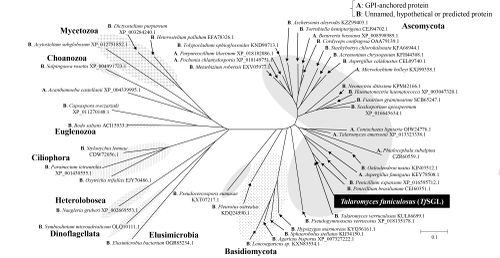
In Glycoside hydrolase family 162, a β-1,2-glucanase only from Talaromyces funiculosus (TfSGL) has been identified, characterized and structurally analyzed as of June 4th, 2019 [1]. The enzyme specifically hydrolyzes both cyclic and linear β-1,2-glucans, which comprise a β-linked glucosyl backbone, and preferably releases sophorose (Glc-β-1,2-Glc) from the reducing end of linear β-1,2-glucan [1]. Almost all of the family members are from Eukaryotes [1]
Kinetics and Mechanism
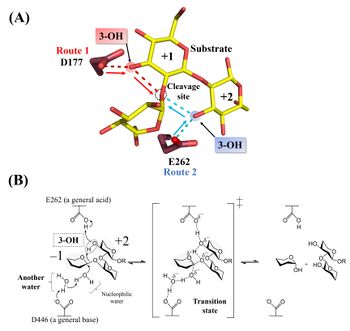
Hydrolysis of cyclic β-1,2-glucan by TfSGL suggests that the enzyme is endo-type [1]. The 1H-NMR analysis of the anomeric configurations of hydrolysates indicates that TfSGL has an inverting mechanism. Analysis of the change of the degree of optical rotation during hydrolysis of β-1,2-glucan also supported this mechanism [1].
Structural analysis (see “Three-dimensional structures”) and mutational analysis suggest that D446 activates the nucleophilic water via another water as a general acid [1]. These analyses also suggest that D177 and/or E262 act as a general acid via the 3-hydroxy groups of the Glc moieties (see below) [1]. According to the action pattern analysis using β-1,2-glucopentaose derivatives deoxygenated at their 3-hydroxy groups in the first or second Glc moiety from the reducing end, E262 was clearly determined to be a general acid. The 3-hydroxy group of the Glc moiety at subsite +2 mediates protonation of glycosidic bond oxygen atom [1]. The reaction mechanism of TfSGL is quite unique in that both reaction pathways involving a general acid and a general base are non-canonical [1].
Catalytic Residues
The general acid and base of TfSGL are E262 and D446, respectively [1]. Both residues are highly conserved in GH162 enzymes. The general acid of TfSGL is well superimposed with an acidic residue in a GH144 bacterial β-1,2-glucanase from Chitinophaga pinensis (CpSGL), whereas the general base is not superimposed [1, 2]. Although the reaction mechanisms of GH144 are unclear as of June 4th, 2019, TfSGL is clearly different from GH144 in the reaction mechanism based on structural comparison [1].
Three-dimensional structures
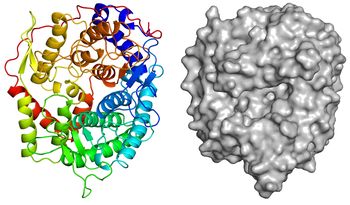
The apo-structure of the recombinant TfSGL (TfSGLr) was determined at 2.0 Å using the iodide single-wavelength anomalous diffraction phasing method (PDB 6IMU) [1]. The overall structure comprises an (α/α)6 toroid fold [1]. The complex structures with sophorose (PDB 6IMV) and the Michaelis complex of an inactive TfSGLr-mutant (E262Q) with a β-1,2-glucoheptaose (PDB 6IMW) were also determined by soaking of crystals in sophorose and β-1,2-glucan, respectively [1]. TfSGLr has a cleft crossing the surface of the structure and there is a large active pocket at the center of the cleft [1]. Interestingly, although TfSGL and GH144 enzymes are quite different in their amino acid sequences, their overall structures and the positions of the substrates in their catalytic pockets are similar [1]. TfSGLr has slight structural similarity to GH15 and GH8 enzymes.
Family Firsts
- First stereochemistry determination
- A fungal β-1,2-glucanase from Talaromyces funiculosus by the NMR analysis and the analysis of the change of the degree of optical rotation [1].
- First general acid residue identification
- A fungal β-1,2-glucanase from Talaromyces funiculosus by the structural analysis, the mutational analysis and the action pattern analysis of β-1,2-sophoropentaose derivatives [1].
- First general base residue identification
- A fungal β-1,2-glucanase from Talaromyces funiculosus by the structural analysis and the mutational analysis [1].
- First 3-D structure
- A fungal β-1,2-glucanase from Talaromyces funiculosus using the iodide single-wavelength anomalous diffraction phasing method [1].
References
- Tanaka N, Nakajima M, Narukawa-Nara M, Matsunaga H, Kamisuki S, Aramasa H, Takahashi Y, Sugimoto N, Abe K, Terada T, Miyanaga A, Yamashita T, Sugawara F, Kamakura T, Komba S, Nakai H, and Taguchi H. (2019). Identification, characterization, and structural analyses of a fungal endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem. 2019;294(19):7942-7965. DOI:10.1074/jbc.RA118.007087 |
- Abe K, Nakajima M, Yamashita T, Matsunaga H, Kamisuki S, Nihira T, Takahashi Y, Sugimoto N, Miyanaga A, Nakai H, Arakawa T, Fushinobu S, and Taguchi H. (2017). Biochemical and structural analyses of a bacterial endo-β-1,2-glucanase reveal a new glycoside hydrolase family. J Biol Chem. 2017;292(18):7487-7506. DOI:10.1074/jbc.M116.762724 |