CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Glycoside Hydrolase Family 23"
| Line 42: | Line 42: | ||
Based on the complexes formed with 1,6-anhydromuropeptide <cite>10</cite> or bulgecin <cite>11</cite>, a substrate-assisted mechanism, analogous to the family GH18 chitinases and chitobiases and family GH20 N-acetyl-β-hexosaminidases, has been invoked for the lytic transglycosyales. Thus, the catalytic Glu73 is proposed to serve initially as an acid catalyst to donate a proton to the glycosidic oxygen of the linkage to be cleaved leading to the formation of an intermediate with oxocarbenium ion character (Figure 2). In the absence of an anion/nucleophile in close proximity to stabilize this oxocarbenium intermediate, the lytic transglycosylases would employ anchimeric assistance of the MurNAc 2-acetamido group resulting in the formation an oxazolinium ion intermediate. This would be followed by abstraction of the C-6 hydroxyl proton of the oxazolinium species involving Glu73 which now serves as the base catalyst leading to nucleophilic attack and the formation of 1,6-anhydromuramic acid product. | Based on the complexes formed with 1,6-anhydromuropeptide <cite>10</cite> or bulgecin <cite>11</cite>, a substrate-assisted mechanism, analogous to the family GH18 chitinases and chitobiases and family GH20 N-acetyl-β-hexosaminidases, has been invoked for the lytic transglycosyales. Thus, the catalytic Glu73 is proposed to serve initially as an acid catalyst to donate a proton to the glycosidic oxygen of the linkage to be cleaved leading to the formation of an intermediate with oxocarbenium ion character (Figure 2). In the absence of an anion/nucleophile in close proximity to stabilize this oxocarbenium intermediate, the lytic transglycosylases would employ anchimeric assistance of the MurNAc 2-acetamido group resulting in the formation an oxazolinium ion intermediate. This would be followed by abstraction of the C-6 hydroxyl proton of the oxazolinium species involving Glu73 which now serves as the base catalyst leading to nucleophilic attack and the formation of 1,6-anhydromuramic acid product. | ||
| − | For the g-type lysozymes, recent studies support a typical inverting mechanism of action <cite>1 2</cite>. In fact, it is possible that two catalytic general base residues, a primary and a secondary residue, position a catalytic water molecule and abstract a proton to | + | For the g-type lysozymes, recent studies support a typical inverting mechanism of action <cite>1 2</cite>. In fact, it is possible that two catalytic general base residues, a primary and a secondary residue, position a catalytic water molecule and abstract a proton to affect substrate hydrolysis by the single displacement mechanism. With GEWL, Asp97 and Asp86, respectively, are proposed to serve this function while these would be represented by the conserved Asp101 and Asp90 residues in the g-type lysozyme from Atlantic cod fish. Indeed, the double replacement of Asp101 and Asp90 in the cod lysozyme results in over a 300-fold decrease in activity <cite>1</cite>. |
== Three-dimensional structures == | == Three-dimensional structures == | ||
| − | Three-dimensional structures are available for several Family GH23 enzymes, the first solved being that of GEWL<cite>6 12</cite>. The catalytic domain of each enzyme possesses the well characterized α+β “lysozyme fold” for avian lysozymes. However, there are distinct structural differences between the two classes of enzymes. Most notably, the environment of the active site in lytic transglycosylases, particularly around the catalytic acid/base, is more hydrophobic compared to that of GEWL. This distinction may account for the difference in the reaction mechanisms of the two enzymes. | + | Three-dimensional structures are available for several Family GH23 enzymes, the first solved being that of GEWL <cite>6 12</cite>. The catalytic domain of each enzyme possesses the well characterized α+β “lysozyme fold” for avian lysozymes. However, there are distinct structural differences between the two classes of enzymes. Most notably, the environment of the active site in lytic transglycosylases, particularly around the catalytic acid/base, is more hydrophobic compared to that of GEWL. This distinction may account for the difference in the reaction mechanisms of the two enzymes. |
Revision as of 12:45, 17 February 2010
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Anthony Clarke^^^
- Responsible Curator: ^^^Anthony Clarke^^^
| Glycoside Hydrolase Family GHnn | |
| Clan | GH-x |
| Mechanism | retaining/inverting |
| Active site residues | known |
| CAZy DB link | |
| http://www.cazy.org/fam/GHnn.html | |
Substrate specificities
The glycoside hydrolases of this family are lytic transglyosylases (also referred to as peptidoglycan lyases) of both bacterial and bacteriophage origin, and family G lysozymes (EC 3.2.1.17; muramidase, peptidoglycan N-acetylmuramoylhydrolase, 1,4-β-N-acetylmuramidase, N-acetylmuramoylhydrolase) of eukaryotic origin. Both of these enzymes are active on peptidoglycan, but only the lysozymes are active on chitin and chitooligosaccharides. No other activities have been observed.
Kinetics and Mechanism
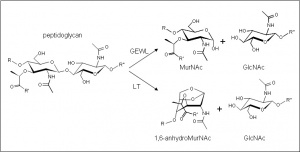
The enzymes of this family cleave the β-1,4 linkage between N-acetylmuramoyl and N-acetylglucosaminyl residues in peptidoglycan (Figure 1) . Only the lysozymes of this family are capable of releasing N-acetyl-d-glucosamine residues from chitodextrins, and neither catalyze (inter) transglycosylation reactions. The mechanism of the family G lysozymes has not been determined experimentally, but theoretical considerations based on crystallographic data [1] and modeling studies [2] suggest that they are inverting enzymes. On the other hand, the lytic transglycosidases, strictly speaking, are retaining enzymes. However, unlike lysozyme, they are not hydrolases but rather catalyse an intramolecular glycosyl transferase reaction onto the C-6 hydroxyl group of the muramoyl residue leading to the generation of a terminal 1,6-anhdyromuramoyl product thus lacking a reducing end [3]. The lytic transglycosylases require the peptide side chains in peptidoglycan for activity, accounting for their inactivity against chitin or chitooligosaccharides [4]. No detailed analyses involving both steady state and pre-steady state kinetic studies have been reported.
Catalytic Residues
Until recently, the family GH23 enzymes were thought to have only a single catalytic residue at their catalytic centre. The identity of the catalytic acid/base residue of the lysozymes was first inferred by X-ray crystallography of goose egg-white lysozyme (GEWL) as Glu 73 [5, 6]. Likewise, analysis of the crystal structure of the soluble lytic transglycosylase 70 (Slt70) from Escherichia coli identified Glu as the lone catalytic residue[7]. Indeed, replacement of each respective residue results in loss of catalytic activity [8, 9]. The mechanism of action of the family GH23 enzymes has yet to be proven experimentally, but examination of crystal structures and theoretical considerations has led to separate proposals for the two classes of enzymes.
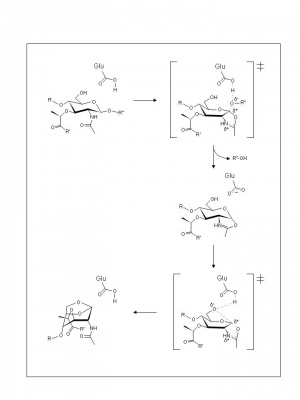
Based on the complexes formed with 1,6-anhydromuropeptide [10] or bulgecin [11], a substrate-assisted mechanism, analogous to the family GH18 chitinases and chitobiases and family GH20 N-acetyl-β-hexosaminidases, has been invoked for the lytic transglycosyales. Thus, the catalytic Glu73 is proposed to serve initially as an acid catalyst to donate a proton to the glycosidic oxygen of the linkage to be cleaved leading to the formation of an intermediate with oxocarbenium ion character (Figure 2). In the absence of an anion/nucleophile in close proximity to stabilize this oxocarbenium intermediate, the lytic transglycosylases would employ anchimeric assistance of the MurNAc 2-acetamido group resulting in the formation an oxazolinium ion intermediate. This would be followed by abstraction of the C-6 hydroxyl proton of the oxazolinium species involving Glu73 which now serves as the base catalyst leading to nucleophilic attack and the formation of 1,6-anhydromuramic acid product.
For the g-type lysozymes, recent studies support a typical inverting mechanism of action [1, 2]. In fact, it is possible that two catalytic general base residues, a primary and a secondary residue, position a catalytic water molecule and abstract a proton to affect substrate hydrolysis by the single displacement mechanism. With GEWL, Asp97 and Asp86, respectively, are proposed to serve this function while these would be represented by the conserved Asp101 and Asp90 residues in the g-type lysozyme from Atlantic cod fish. Indeed, the double replacement of Asp101 and Asp90 in the cod lysozyme results in over a 300-fold decrease in activity [1].
Three-dimensional structures
Three-dimensional structures are available for several Family GH23 enzymes, the first solved being that of GEWL [6, 12]. The catalytic domain of each enzyme possesses the well characterized α+β “lysozyme fold” for avian lysozymes. However, there are distinct structural differences between the two classes of enzymes. Most notably, the environment of the active site in lytic transglycosylases, particularly around the catalytic acid/base, is more hydrophobic compared to that of GEWL. This distinction may account for the difference in the reaction mechanisms of the two enzymes.
Family Firsts
- First identification of lytic transglycosylase
- Soluble lytic transglycosylase 70 [3].
- First catalytic nucleophile identification
- For g-type lysozymes [2]; Not applicable for lytic transglycosylases.
- First general acid/base residue identification
- Inferred by X-ray crystallography of goose egg-white lysozyme [6].
- First 3-D structure
- Goose egg-white lysozyme [12].
References
- Helland R, Larsen RL, Finstad S, Kyomuhendo P, and Larsen AN. (2009). Crystal structures of g-type lysozyme from Atlantic cod shed new light on substrate binding and the catalytic mechanism. Cell Mol Life Sci. 2009;66(15):2585-98. DOI:10.1007/s00018-009-0063-x |
- Hirakawa H, Ochi A, Kawahara Y, Kawamura S, Torikata T, and Kuhara S. (2008). Catalytic reaction mechanism of goose egg-white lysozyme by molecular modelling of enzyme-substrate complex. J Biochem. 2008;144(6):753-61. DOI:10.1093/jb/mvn133 |
- Höltje JV, Mirelman D, Sharon N, and Schwarz U. (1975). Novel type of murein transglycosylase in Escherichia coli. J Bacteriol. 1975;124(3):1067-76. DOI:10.1128/jb.124.3.1067-1076.1975 |
- Romeis T, Vollmer W, and Höltje JV. (1993). Characterization of three different lytic transglycosylases in Escherichia coli. FEMS Microbiol Lett. 1993;111(2-3):141-6. DOI:10.1111/j.1574-6968.1993.tb06376.x |
- Weaver LH, Grütter MG, and Matthews BW. (1995). The refined structures of goose lysozyme and its complex with a bound trisaccharide show that the "goose-type" lysozymes lack a catalytic aspartate residue. J Mol Biol. 1995;245(1):54-68. DOI:10.1016/s0022-2836(95)80038-7 |
- Weaver LH, Grütter MG, Remington SJ, Gray TM, Isaacs NW, and Matthews BW. (1984-1985). Comparison of goose-type, chicken-type, and phage-type lysozymes illustrates the changes that occur in both amino acid sequence and three-dimensional structure during evolution. J Mol Evol. 1984-1985;21(2):97-111. DOI:10.1007/BF02100084 |
- van Asselt EJ, Thunnissen AM, and Dijkstra BW. (1999). High resolution crystal structures of the Escherichia coli lytic transglycosylase Slt70 and its complex with a peptidoglycan fragment. J Mol Biol. 1999;291(4):877-98. DOI:10.1006/jmbi.1999.3013 |
- van Asselt EJ, Dijkstra AJ, Kalk KH, Takacs B, Keck W, and Dijkstra BW. (1999). Crystal structure of Escherichia coli lytic transglycosylase Slt35 reveals a lysozyme-like catalytic domain with an EF-hand. Structure. 1999;7(10):1167-80. DOI:10.1016/s0969-2126(00)80051-9 |
- Kuroki R, Weaver LH, and Matthews BW. (1999). Structural basis of the conversion of T4 lysozyme into a transglycosidase by reengineering the active site. Proc Natl Acad Sci U S A. 1999;96(16):8949-54. DOI:10.1073/pnas.96.16.8949 |
- Thunnissen AM, Rozeboom HJ, Kalk KH, and Dijkstra BW. (1995). Structure of the 70-kDa soluble lytic transglycosylase complexed with bulgecin A. Implications for the enzymatic mechanism. Biochemistry. 1995;34(39):12729-37. DOI:10.1021/bi00039a032 |
- Thunnissen AM, Isaacs NW, and Dijkstra BW. (1995). The catalytic domain of a bacterial lytic transglycosylase defines a novel class of lysozymes. Proteins. 1995;22(3):245-58. DOI:10.1002/prot.340220305 |
- Grütter MG, Weaver LH, and Matthews BW. (1983). Goose lysozyme structure: an evolutionary link between hen and bacteriophage lysozymes?. Nature. 1983;303(5920):828-31. DOI:10.1038/303828a0 |