CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Auxiliary Activity Family 3
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
- Author: ^^^Roland Ludwig^^^ and ^^^Daniel Kracher^^^
- Responsible Curator: ^^^Roland Ludwig^^^
| Auxiliary Activity Family AA3 | |
| Clan | AA3 |
| Mechanism | FAD-dependent substrate oxidation |
| Active site residues | known |
| CAZy DB link | |
| https://www.cazy.org/AA3.html | |
General properties and substrate specificities
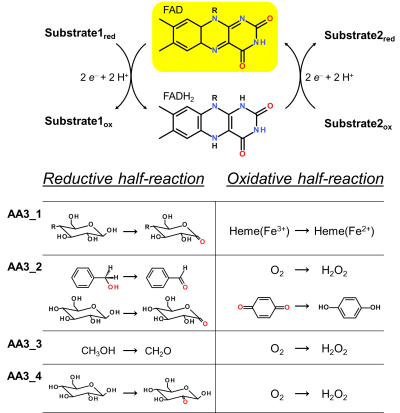
Enzymes of the CAZY family AA3 are widespread and catalyse the oxidation of alcohols or carbohydrates with the concomitant formation of hydrogen peroxide or hydroquinones [1]. AA3 enzymes are most abundant in wood-degrading fungi where they typically display a high multigenicity [2]. The main function of fungal AA3 enzymes is the stimulation of lignocellulose degradation in cooperation with other AA-enzymes such as peroxidases (AA2) [3] or lytic polysaccharide monooxygenases (AA9) [4, 5]. In yeast, AA3 enzymes are involved in the catabolism of alcohols [6], and some AA3 genes identified in insects are thought to be relevant for immunity and development [7, 8]. Recently, a bacterial AA3 enzyme with unknown biological function was isolated and characterized [9]. The functionally diverse enzymes of family AA3 all belong to the structurally related glucose-methanol-choline (GMC) family of oxidoreductases and require a flavin-adenine dinucleotide (FAD) cofactor for catalytic activity. Based on their sequences members of the AA3 family were divided into four subfamilies in the CAZy database (Figure 1). Family AA3_1 contains the flavodehydrogenase domain of cellobiose dehydrogenase (EC 1.1.99.18), family AA3_2 includes aryl alcohol oxidase (EC 1.1.3.7), glucose oxidase (EC 1.1.3.4), glucose dehydrogenase (EC 1.1.5.9) and pyranose dehydrogenase (EC 1.1.99.29), family AA3_3 consists of alcohol (methanol) oxidases (EC 1.1.3.13) and family AA3_4 comprises pyranose oxidoreductases (EC 1.1.3.10).
Subfamily AA3_1: cellobiose dehydrogenase
Cellobiose dehydrogenases (CDHs) are extracellular flavocytochromes that were first described in 1974 [10]. CDHs are exclusively found in wood-degrading and phytopathogenic fungi belonging to the phyla Basidiomycota (Class-I CDHs) and Ascomycota (Class-II and -III CDHs) [11]. They oxidize a wide variety of lignocellulose-derived saccharides to their corresponding sugar lactones. CDHs show a high preference for soluble, β-(1,4)-interlinked saccharides and scarcely oxidize monosaccharides. A common feature of all CDHs is their complex bipartite structure, which comprises a C-terminal cytochrome-binding domain (CYT) and a larger, catalytic flavodehydrogenase (DH) domain encoded within a single polypeptide chain [12] (Figure 2A). Both domains are connected by a flexible linker which typically comprises 15 – 30 amino acids. An important in vivo function of CDH is the reduction and activation of family AA9 lytic polysaccharide monooxygenases via its heme b domain [13, 14]. Recently, the holoenzyme structures of Neurospora crassa CDH (pdb: 4qi7) and Myriococcum thermophilum CDH (pdb: 4qi5) were reported. Two structures of N. crassa CDH showed an “open-state” conformation in which DH and CYT were spatially separated, whereas a structure of M. thermophilum CDH showed a “closed-state” conformation in which the propionate arm of the cytochrome domain interacted with the catalytic centre in DH [14]. Analysis by small angle scattering also suggested a number of possible intermediate conformers that exist in solution [14, 15]. While the closed-state allows interdomain electron transfer from DH to CYT, reduction of electron acceptors (e.g. AA9 enzymes) might occur in the open-state, in which the heme cofactor is fully accessible.
Subfamily AA3_2
Aryl Alcohol Oxidase/Dehydrogenase
Aryl alcohol oxidoreductases (AAO) are secreted by basidiomycete fungi and proposed to play a crucial role in lignin degradation. The X-ray crystal structure of the most widely studied AAO from Pleurotus eryngii (pdb: 3fim) showed a non-covalently bound FAD cofactor in the active-site [16] (Figure 2C). Access to the active site is restricted by three aromatic residues, which interact with both the alcohol substrate and oxygen. The substrate scope of AAO includes secreted benzyl alcohols from the secondary metabolism (e.g. veratryl alcohol), or related alcohols that accumulate during lignin degradation [3]. During the reductive half-reaction, the primary alcohol of the substrate is two-electron oxidized to form the corresponding aldehyde. In addition, aromatic aldehydes can undergo further oxidation yielding the corresponding acids. The concomitantly formed hydrogen peroxide is considered essential for the activity of lignin degrading peroxidases. Recently, aryl-alcohol quinone oxidoreductases (AAQO) with a high sequence identity to Pleurotus eryngii AAO were identified in the genome of the basidiomycete Pycnoporus cinnabarinus [17]. While these enzymes showed a substrate spectrum similar to known AAOs, oxygen reduction was insignificant or very low relative to other characterised AAOs. However, reoxidation of FADH2 was very efficient with benzoquinone or phenoxy-radicals, which are formed by fungal laccases, suggesting a different functional role of AAQOs.
Subfamily 3 - GOx
Kinetics and Mechanism
Glucose oxidase (GOx, syn. GOD, EC 1.1.x.x., glucose:oxygen 1-oxidoreductase) catalyzes the regioselective oxidation of β-D-glucose to gluconic acid by utilizing molecular oxygen as an electron acceptor with simultaneous production of hydrogen peroxide. Glucose oxidases are produced by a number of fungi and insects. Proposed functions are related to the peroxide-producing abilities of the enzyme and include the preservation of honey, microbial defense as well as the support of H2O2 dependent ligninases in wood degrading fungi [18].
In the oxidative half reaction of GOx, two-electron oxidation of D-glucose at C1 results in the formation of gluconolactone and reduced cofactor (FADH2). Reduction by glucose occurs via a hydride ion transfer to the N-5 position of the FAD. Oxygen reduction to H2O2 involves the uptake of two electrons (from FADH2) and two protons. A highly conserved His (His516 in Aspergillus niger GOx) in the active site of the enzyme was identified by site directed mutagenesis as the catalytic base [19].
Catalytic Residues
Content is to be added here.
Three-dimensional structures
Content is to be added here.
Family Firsts
- First GOx identified
- Glucose oxidase from Aspergillus niger in 1928 [20].
- First 3D structure
- Glucose oxidase from Aspergillus niger (1GAL; [21]).
References
- Levasseur A, Drula E, Lombard V, Coutinho PM, and Henrissat B. (2013). Expansion of the enzymatic repertoire of the CAZy database to integrate auxiliary redox enzymes. Biotechnol Biofuels. 2013;6(1):41. DOI:10.1186/1754-6834-6-41 |
- Sützl L, Laurent CVFP, Abrera AT, Schütz G, Ludwig R, and Haltrich D. (2018). Multiplicity of enzymatic functions in the CAZy AA3 family. Appl Microbiol Biotechnol. 2018;102(6):2477-2492. DOI:10.1007/s00253-018-8784-0 |
- Hernández-Ortega A, Ferreira P, and Martínez AT. (2012). Fungal aryl-alcohol oxidase: a peroxide-producing flavoenzyme involved in lignin degradation. Appl Microbiol Biotechnol. 2012;93(4):1395-410. DOI:10.1007/s00253-011-3836-8 |
- Kracher D, Scheiblbrandner S, Felice AK, Breslmayr E, Preims M, Ludwicka K, Haltrich D, Eijsink VG, and Ludwig R. (2016). Extracellular electron transfer systems fuel cellulose oxidative degradation. Science. 2016;352(6289):1098-101. DOI:10.1126/science.aaf3165 |
- Garajova S, Mathieu Y, Beccia MR, Bennati-Granier C, Biaso F, Fanuel M, Ropartz D, Guigliarelli B, Record E, Rogniaux H, Henrissat B, and Berrin JG. (2016). Single-domain flavoenzymes trigger lytic polysaccharide monooxygenases for oxidative degradation of cellulose. Sci Rep. 2016;6:28276. DOI:10.1038/srep28276 |
- Goswami P, Chinnadayyala SS, Chakraborty M, Kumar AK, and Kakoti A. (2013). An overview on alcohol oxidases and their potential applications. Appl Microbiol Biotechnol. 2013;97(10):4259-75. DOI:10.1007/s00253-013-4842-9 |
- Iida K, Cox-Foster DL, Yang X, Ko WY, and Cavener DR. (2007). Expansion and evolution of insect GMC oxidoreductases. BMC Evol Biol. 2007;7:75. DOI:10.1186/1471-2148-7-75 |
- Sun W, Shen YH, Yang WJ, Cao YF, Xiang ZH, and Zhang Z. (2012). Expansion of the silkworm GMC oxidoreductase genes is associated with immunity. Insect Biochem Mol Biol. 2012;42(12):935-45. DOI:10.1016/j.ibmb.2012.09.006 |
-
Mendes S, Banha C, Madeira J, Santos D, Miranda V, Manzanera M, Ventura MR, van Berkel WJH, Martins LO. Characterization of a bacterial pyranose 2-oxidase from Arthrobacter siccitolerans. 2016 J Mol Catal B Enzym 133, S34–S43
-
Westermark U, Eriksson KE. Cellobiose:Quinone Oxidoreductase, a New Wood-degrading Enzyme from White-rot Fungi. 1974 Acta Chem Scand 1974 28b:209–214.
-
Kracher D, Ludwig R. Cellobiose dehydrogenase: An essential enzyme for lignocellulose degradation in nature - A review. 2016 Bodenkultur 67:145–163.
- Hallberg BM, Henriksson G, Pettersson G, Vasella A, and Divne C. (2003). Mechanism of the reductive half-reaction in cellobiose dehydrogenase. J Biol Chem. 2003;278(9):7160-6. DOI:10.1074/jbc.M210961200 |
- Phillips CM, Beeson WT, Cate JH, and Marletta MA. (2011). Cellobiose dehydrogenase and a copper-dependent polysaccharide monooxygenase potentiate cellulose degradation by Neurospora crassa. ACS Chem Biol. 2011;6(12):1399-406. DOI:10.1021/cb200351y |
- Tan TC, Kracher D, Gandini R, Sygmund C, Kittl R, Haltrich D, Hällberg BM, Ludwig R, and Divne C. (2015). Structural basis for cellobiose dehydrogenase action during oxidative cellulose degradation. Nat Commun. 2015;6:7542. DOI:10.1038/ncomms8542 |
- Bodenheimer AM, O'Dell WB, Oliver RC, Qian S, Stanley CB, and Meilleur F. (2018). Structural investigation of cellobiose dehydrogenase IIA: Insights from small angle scattering into intra- and intermolecular electron transfer mechanisms. Biochim Biophys Acta Gen Subj. 2018;1862(4):1031-1039. DOI:10.1016/j.bbagen.2018.01.016 |