CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Difference between revisions of "Carbohydrate Binding Module Family 102"
(Created page with " <!-- RESPONSIBLE CURATORS: Please replace the {{UnderConstruction}} tag below with {{CuratorApproved}} when the page is ready for wider public consumption --> {{UnderConstruc...") |
|||
| Line 18: | Line 18: | ||
== Ligand specificities == | == Ligand specificities == | ||
| − | + | The dissected C-terminal CBM102 of a surface glycan-binding protein (SGBP; containing four CBM102s) of ''Christiangramia forsetii'' KT0803<sup>T</sup> is binding laminarin (from ''Laminaria digitata'') and laminarin-derived oligosaccharides, which was confirmed by affinity gel electrophoresis and/or isothermal titration calorimetry <cite>Zuehlke2024</cite>. The CBM102 had a higher affinity to laminarin (K<sub>a</sub> ~1.04 x 10<sup>5</sup> M<sup>-1</sup>) than to laminariheptaose (K<sub>a</sub> ~1.87 x 10<sup>4</sup> M<sup>-1</sup>) and laminaripentaose (K<sub>a</sub> ~1.39 x 10<sup>4</sup> M<sup>-1</sup>). Laminaribiose was not binding. | |
| − | + | In addition, the dissected C-terminal CBM102 had a higher affinity to laminarin than two CBM102 tandem constructs of the SGBP (K<sub>a</sub> ~2.76 x 10<sup>4</sup> M<sup>-1</sup> or ~3.63 x 10<sup>4</sup> M<sup>-1</sup>) and the full length SGBP (K<sub>a</sub> ~3.09 x 10<sup>4</sup> M<sup>-1</sup>) <cite>Zuehlke2024</cite>. | |
== Structural Features == | == Structural Features == | ||
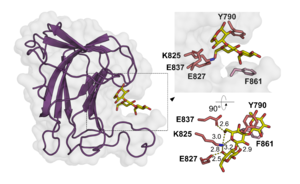
| − | + | The CBM102 displays a β-sandwich fold established by two antiparallel β-sheets forming a convex and a concave face. The 3D X-ray crystal structure of the C-terminal CBM102 of the ''C. forsetii'' SGBP showed that together with two expansive loops, the concave face provides a narrow cleft to accommodate laminaritriose <cite>Zuehlke2024</cite>. CBM102 thus represents a [[Carbohydrate-binding_modules|type B CBM]]. Three residues located in the concave face mediate polar interactions with laminaritriose (Lys825, Glu827 and Glu837), while one aromatic residue of each loop (Tyr790, Phe861) is involved in CH-π interaction with the ligand. However, the ''C. forsetii'' protein contains three additional CBM102s. Alphafold2 <cite>Mirdita2022</cite> predictions showed that two of them are highly similar to the 3D crystal structure of the C-terminal CBM102, but that one CBM102 is missing one of the loops shaping the pocket, which results in a very shallow cleft. This was speculated to represent an adaptation to highly branched laminarins <cite>Zuehlke2024</cite>. | |
| − | + | [[File:Fig.1.png|thumb|3D crystal structure of C-terminal CBM102 of the ''C. forsetii'' SGBP with laminaritriose. Highlighted are interacting residues.]] | |
| − | |||
| − | |||
| + | [[File:Fig.2.png|thumb|Surfaces of the four CBM102s of the ''C. forsetii'' SGBP. Left: 3D crystal structure of the C-terminal CBM102 (IV) with laminaritriose. Right: Alphafold2-predicted structures (I-III). Module III is missing one of the loops, which form the groove.]] | ||
== Functionalities == | == Functionalities == | ||
| − | '' | + | The CBM102-containing SGBP is tethered to the outer membrane of ''C. forsetii'' and is abundantly produced in laminarin-grown cells <cite>Kabisch2014</cite>. With four CBM102s, the SGBP provides multiple laminarin docking sites <cite>Zuehlke2024</cite>. The protein further contains two N-terminal Ig-like domains that precede the CBM102s, which presumably act as spacers to ensure distance to the membrane and exposure of binding sites as suggested before for β-glucan binding SGBPs <cite>Tamura2021</cite>. The multiplicity of CBM102 in the ''C. forsetii'' protein was suggested to render the SGBP an optimal 'sugar-trapping antenna' on the bacterial surface, which was supported by an elongated protein shape as determined by small angle X ray scattering <cite>Zuehlke2024</cite>. However, isothermal titration calorimetry showed that two laminarin molecules were binding at maximum and that affinity was not increased by multiple CBM102s. In bacterial metagenome-assembled genomes from phytoplankton blooms in the North Sea, CBM102 was detected together with catalytic modules, indicating also other functions (increased catalysis) <cite>Zuehlke2024</cite>. The majority of associated catalytic modules were GH16_3s, but also rare examples of GH2, GH5_34, GH81, GH162. Within this dataset, CBM102 also co-occurred with CBM4 and CBM6. CBM102-only-proteins contained up to eight repeats of CBM102. The production of CBM102-containing proteins correlated with the course of the phytoplankton bloom, which underlines the relevance of CBM102 in marine β-glucan use. |
| − | |||
| − | |||
| − | |||
== Family Firsts == | == Family Firsts == | ||
;First Identified | ;First Identified | ||
| − | + | CBM102 was first identified in an SGBP encoded in a laminarin utilization locus of ''Christiangramia forsetii'' KT0803<sup>T</sup> <cite>Zuehlke2024</cite>, a marine flavobacterium isolated from surface waters in the North Sea <cite>Eilers2001</cite>. 362 CBM102-containing sequences were detected in bacterial metagenome-assembled genomes (555 representative MAGs of which 201 belonged to ''Bacteroidota'') retrieved from bacterial biomass from phytoplankton blooms of three respective years (2016, 2018 and 2020) in the North Sea. Sequences mostly belonged to ''Bacteroidota'', but also to ''Proteobacteria'', ''Myxococcota'' and ''Actinobacteriota'' <cite>Zuehlke2024</cite>. The association with GHs (mostly GH16_3) within this dataset led to the creation of the CBM family 102. | |
;First Structural Characterization | ;First Structural Characterization | ||
| − | + | The C-terminal CBM102 of the laminarin PUL-encoded SGBP of ''C. forsetii'' KT0803<sup>T</sup> is the first structurally characterized member of the family (PDB ID [https://www.rcsb.org/structure/unreleased/8QX6 8QX6]). The 3D crystal structure was obtained in complex with laminaritriose. | |
== References == | == References == | ||
<biblio> | <biblio> | ||
| − | # | + | #Zuehlke2024 pmid=38757353 |
| − | # | + | #Mirdita2022 pmid=35637307 |
| − | + | #Kabisch2014 pmid=24522261 | |
| − | + | #Tamura2021 pmid=33587952 | |
| − | # | + | #Eilers2001 pmid=11679337 |
| − | # | ||
| − | # | ||
</biblio> | </biblio> | ||
Revision as of 06:49, 6 September 2024
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
| CAZy DB link | |
| https://www.cazy.org/CBM102.html |
Ligand specificities
The dissected C-terminal CBM102 of a surface glycan-binding protein (SGBP; containing four CBM102s) of Christiangramia forsetii KT0803T is binding laminarin (from Laminaria digitata) and laminarin-derived oligosaccharides, which was confirmed by affinity gel electrophoresis and/or isothermal titration calorimetry [1]. The CBM102 had a higher affinity to laminarin (Ka ~1.04 x 105 M-1) than to laminariheptaose (Ka ~1.87 x 104 M-1) and laminaripentaose (Ka ~1.39 x 104 M-1). Laminaribiose was not binding.
In addition, the dissected C-terminal CBM102 had a higher affinity to laminarin than two CBM102 tandem constructs of the SGBP (Ka ~2.76 x 104 M-1 or ~3.63 x 104 M-1) and the full length SGBP (Ka ~3.09 x 104 M-1) [1].
Structural Features
The CBM102 displays a β-sandwich fold established by two antiparallel β-sheets forming a convex and a concave face. The 3D X-ray crystal structure of the C-terminal CBM102 of the C. forsetii SGBP showed that together with two expansive loops, the concave face provides a narrow cleft to accommodate laminaritriose [1]. CBM102 thus represents a type B CBM. Three residues located in the concave face mediate polar interactions with laminaritriose (Lys825, Glu827 and Glu837), while one aromatic residue of each loop (Tyr790, Phe861) is involved in CH-π interaction with the ligand. However, the C. forsetii protein contains three additional CBM102s. Alphafold2 [2] predictions showed that two of them are highly similar to the 3D crystal structure of the C-terminal CBM102, but that one CBM102 is missing one of the loops shaping the pocket, which results in a very shallow cleft. This was speculated to represent an adaptation to highly branched laminarins [1].
Functionalities
The CBM102-containing SGBP is tethered to the outer membrane of C. forsetii and is abundantly produced in laminarin-grown cells [3]. With four CBM102s, the SGBP provides multiple laminarin docking sites [1]. The protein further contains two N-terminal Ig-like domains that precede the CBM102s, which presumably act as spacers to ensure distance to the membrane and exposure of binding sites as suggested before for β-glucan binding SGBPs [4]. The multiplicity of CBM102 in the C. forsetii protein was suggested to render the SGBP an optimal 'sugar-trapping antenna' on the bacterial surface, which was supported by an elongated protein shape as determined by small angle X ray scattering [1]. However, isothermal titration calorimetry showed that two laminarin molecules were binding at maximum and that affinity was not increased by multiple CBM102s. In bacterial metagenome-assembled genomes from phytoplankton blooms in the North Sea, CBM102 was detected together with catalytic modules, indicating also other functions (increased catalysis) [1]. The majority of associated catalytic modules were GH16_3s, but also rare examples of GH2, GH5_34, GH81, GH162. Within this dataset, CBM102 also co-occurred with CBM4 and CBM6. CBM102-only-proteins contained up to eight repeats of CBM102. The production of CBM102-containing proteins correlated with the course of the phytoplankton bloom, which underlines the relevance of CBM102 in marine β-glucan use.
Family Firsts
- First Identified
CBM102 was first identified in an SGBP encoded in a laminarin utilization locus of Christiangramia forsetii KT0803T [1], a marine flavobacterium isolated from surface waters in the North Sea [5]. 362 CBM102-containing sequences were detected in bacterial metagenome-assembled genomes (555 representative MAGs of which 201 belonged to Bacteroidota) retrieved from bacterial biomass from phytoplankton blooms of three respective years (2016, 2018 and 2020) in the North Sea. Sequences mostly belonged to Bacteroidota, but also to Proteobacteria, Myxococcota and Actinobacteriota [1]. The association with GHs (mostly GH16_3) within this dataset led to the creation of the CBM family 102.
- First Structural Characterization
The C-terminal CBM102 of the laminarin PUL-encoded SGBP of C. forsetii KT0803T is the first structurally characterized member of the family (PDB ID 8QX6). The 3D crystal structure was obtained in complex with laminaritriose.
References
- Zühlke MK, Ficko-Blean E, Bartosik D, Terrapon N, Jeudy A, Jam M, Wang F, Welsch N, Dürwald A, Martin LT, Larocque R, Jouanneau D, Eisenack T, Thomas F, Trautwein-Schult A, Teeling H, Becher D, Schweder T, and Czjzek M. (2024). Unveiling the role of novel carbohydrate-binding modules in laminarin interaction of multimodular proteins from marine Bacteroidota during phytoplankton blooms. Environ Microbiol. 2024;26(5):e16624. DOI:10.1111/1462-2920.16624 |
- Mirdita M, Schütze K, Moriwaki Y, Heo L, Ovchinnikov S, and Steinegger M. (2022). ColabFold: making protein folding accessible to all. Nat Methods. 2022;19(6):679-682. DOI:10.1038/s41592-022-01488-1 |
- Kabisch A, Otto A, König S, Becher D, Albrecht D, Schüler M, Teeling H, Amann RI, and Schweder T. (2014). Functional characterization of polysaccharide utilization loci in the marine Bacteroidetes 'Gramella forsetii' KT0803. ISME J. 2014;8(7):1492-502. DOI:10.1038/ismej.2014.4 |
- Tamura K, Dejean G, Van Petegem F, and Brumer H. (2021). Distinct protein architectures mediate species-specific beta-glucan binding and metabolism in the human gut microbiota. J Biol Chem. 2021;296:100415. DOI:10.1016/j.jbc.2021.100415 |
- Eilers H, Pernthaler J, Peplies J, Glöckner FO, Gerdts G, and Amann R. (2001). Isolation of novel pelagic bacteria from the German bight and their seasonal contributions to surface picoplankton. Appl Environ Microbiol. 2001;67(11):5134-42. DOI:10.1128/AEM.67.11.5134-5142.2001 |