CAZypedia celebrates the life of Senior Curator Emeritus Harry Gilbert, a true giant in the field, who passed away in September 2025.
CAZypedia needs your help!
We have many unassigned pages in need of Authors and Responsible Curators. See a page that's out-of-date and just needs a touch-up? - You are also welcome to become a CAZypedian. Here's how.
Scientists at all career stages, including students, are welcome to contribute.
Learn more about CAZypedia's misson here and in this article. Totally new to the CAZy classification? Read this first.
Glycosyltransferase Family 138
This page is currently under construction. This means that the Responsible Curator has deemed that the page's content is not quite up to CAZypedia's standards for full public consumption. All information should be considered to be under revision and may be subject to major changes.
| Glycosyltransferase Family GT138 | |
| Clan | Fido fold |
| Mechanism | Inverting |
| Active site residues | Known |
| CAZy DB link | |
| https://www.cazy.org/GT138.html | |
Substrate specificities
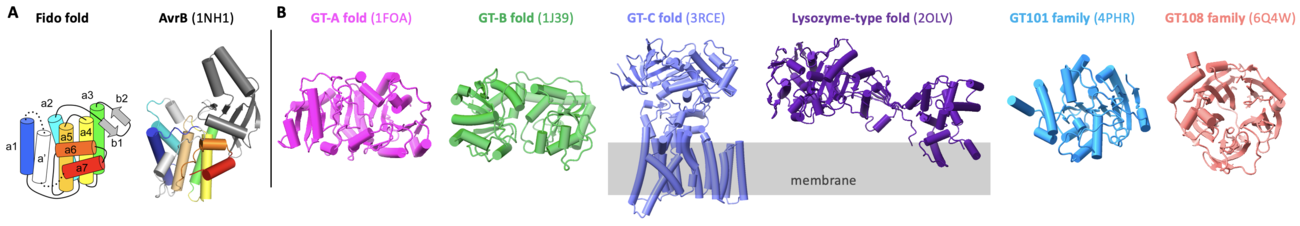
GT138 family of glycosyltransferase is exemplified by AvrB [1]. As a bacterial effector from the plant pathogen Pseudomonas syringae, AvrB utilizes host UDP-rhamnose (or dTDP-rhamnose in vitro) as a co-substrate to modify the host protein RIN4 and causes the programmed cell death (namely hypersensitive response). AvrB contains a Fido domain [2] (Fig. 1A), different from other known glycosyltransferases containing folds of GT-A, GT-B, GT-C, lysozyme-type, GT101, and GT108 (Fig. 1B). Interestingly, Fido proteins can also be enzymes with activities of AMPylation [3], phosphorylation [4], UMPylation [5], and phosphocholination [6, 7]. Therefore, AvrB is a unique Fido protein that functions as a glycosyltransferase.
Kinetics and Mechanism
Content is to be added here.
Catalytic Residues
Content is to be added here.
Three-dimensional structures
AvrB represents the prototype for glycosyltransferases of Fido fold. AvrB contains a relatively large internal domain between helix α2 and helix α3 (Fig. 1A). Other Fido enzyme structures have been determined, though they have diverse activities as mentioned above. These Fido proteins share a similar fold while the primary sequences are divergent.
Family Firsts
- First stereochemistry determination
- Content is to be added here.
- First catalytic nucleophile identification
- Content is to be added here.
- First general acid/base residue identification
- Content is to be added here.
- First 3-D structure
- Content is to be added here.
References
- Peng W, Garcia N, Servage KA, Kohler JJ, Ready JM, Tomchick DR, Fernandez J, and Orth K. (2024). Pseudomonas effector AvrB is a glycosyltransferase that rhamnosylates plant guardee protein RIN4. Sci Adv. 2024;10(7):eadd5108. DOI:10.1126/sciadv.add5108 |
- Kinch LN, Yarbrough ML, Orth K, and Grishin NV. (2009). Fido, a novel AMPylation domain common to fic, doc, and AvrB. PLoS One. 2009;4(6):e5818. DOI:10.1371/journal.pone.0005818 |
- Yarbrough ML, Li Y, Kinch LN, Grishin NV, Ball HL, and Orth K. (2009). AMPylation of Rho GTPases by Vibrio VopS disrupts effector binding and downstream signaling. Science. 2009;323(5911):269-72. DOI:10.1126/science.1166382 |
- Castro-Roa D, Garcia-Pino A, De Gieter S, van Nuland NAJ, Loris R, and Zenkin N. (2013). The Fic protein Doc uses an inverted substrate to phosphorylate and inactivate EF-Tu. Nat Chem Biol. 2013;9(12):811-7. DOI:10.1038/nchembio.1364 |
- Feng F, Yang F, Rong W, Wu X, Zhang J, Chen S, He C, and Zhou JM. (2012). A Xanthomonas uridine 5'-monophosphate transferase inhibits plant immune kinases. Nature. 2012;485(7396):114-8. DOI:10.1038/nature10962 |
- Mukherjee S, Liu X, Arasaki K, McDonough J, Galán JE, and Roy CR. (2011). Modulation of Rab GTPase function by a protein phosphocholine transferase. Nature. 2011;477(7362):103-6. DOI:10.1038/nature10335 |
- Campanacci V, Mukherjee S, Roy CR, and Cherfils J. (2013). Structure of the Legionella effector AnkX reveals the mechanism of phosphocholine transfer by the FIC domain. EMBO J. 2013;32(10):1469-77. DOI:10.1038/emboj.2013.82 |